Deep learning takes fluorescence microscopy into super resolution
UCLA-led team produces images on a laptop that match the quality of those from high-end equipment.
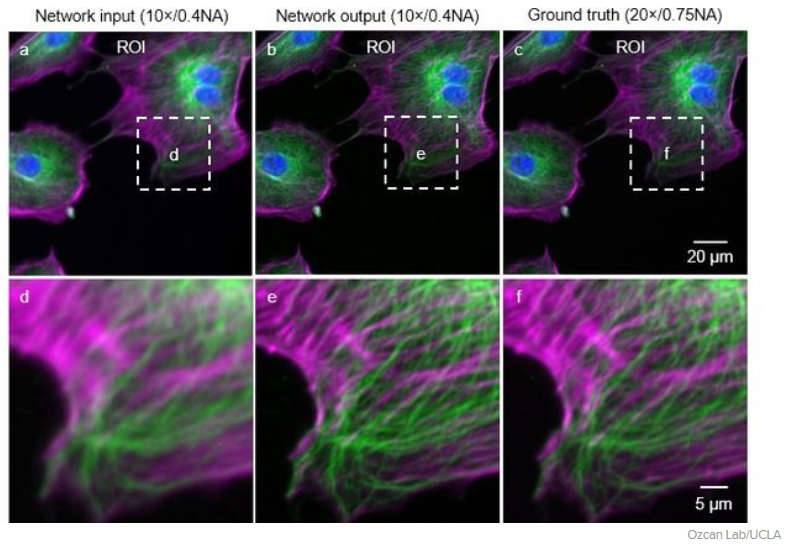
The technique transforms low-resolution images from a fluorescence microscope (a) into super-resolution images (b) that compare favorably with those from high-resolution equipment (c). Images on the bottom row are closeups of those on the top row.
Scientists studying the mysteries of life sometimes rely upon fluorescence microscopy to get a close look at living cells. The technique involves dyeing parts of cells so that they glow under special lighting, revealing cellular structures that measure smaller than one-millionth of a meter.
However, even high-resolution fluorescence microscopes have a hard limit to the amount of detail they can show. Within the past few decades, methods that yield “super-resolution” images have broken that barrier, revealing details at the sub-cellular level, even smaller than one ten-millionth of a meter — an advance that won the 2014 Nobel Prize in Chemistry.
But those strategies come with their own drawbacks: They can be expensive and complex, and they sometimes involve high-intensity light that is toxic to the cells being studied.
Now, UCLA researchers have created a new technique that uses deep learning — a type of artificial intelligence in which machines “learn” through data patterns — to transform lower-resolution fluorescence microscopy images into super resolution. The framework takes images from a simple, inexpensive microscope and produces images that mimic those from more advanced and expensive ones.
The researchers were led by Aydogan Ozcan, UCLA Chancellor’s Professor of Electrical and Computer Engineering and associate director of the California NanoSystems Institute at UCLA; Hongda Wang, a UCLA graduate student, and Yair Rivenson, a UCLA postdoctoral scholar, are the study’s co-first authors. The findings were published online by Nature Methods.
“We need better microscopes to enable discovery at the micro- and nanoscale and allow us to make observations that are otherwise impossible,” Ozcan said, adding that the technology could be an inexpensive and easy-to-use solution for scientists who are researching the molecular workings of cells and other microscopic systems, but who lack the resources to purchase or use more sophisticated equipment.
The scientists’ work could make advanced microscopy more readily accessible to researchers and open paths of discovery throughout science and engineering.
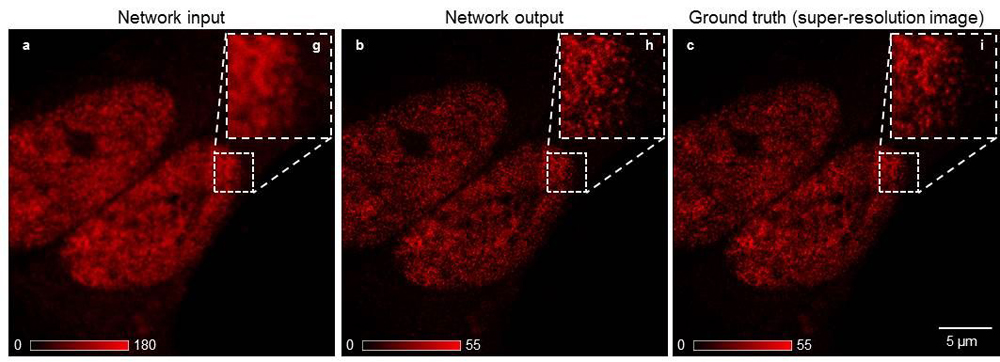
During the experiments, the researchers fed a computer thousands of images of cells and other microscopic structures, taken by five types of fluorescence microscopes. The images were presented in matched pairs, with the object shown in lower resolution and super resolution.
To learn from those images, the system uses a “generative adversarial network,” a model for artificial intelligence in which two algorithms compete. One algorithm tries to create computer-generated super-resolution images from a low-resolution input image, while the second algorithm tries to differentiate between those computer-generated images and existing super-resolution images that are obtained from advanced microscopes.
That “training” needs to be done only once for each type of subject the system needs to learn. After that, the network can improve a low-resolution image it has never “seen” before to match the image resolution from a super-resolution microscope, which eliminates the need for an expensive, high-resolution microscope.
In the study, the UCLA-developed system successfully enhanced the resolution, contrast and depth of field of original images, which were of cell and tissue samples.
“Using a super-resolution microscope requires precise technical skills and expertise,” said Laurent Bentolila, a co-author of the study and the director of the Advanced Light Microscopy/Spectroscopy Laboratory at CNSI. “Seeing that you can now get the same results using deep learning without an advanced and delicate instrument is truly amazing.”
The new approach avoids some of the disadvantages of other super-resolution techniques. For instance, scientists do not need to illuminate the sample with intense light, which can alter cells’ behavior, or even damage or kill them. In addition, it improves resolution based only upon image data. In the study, this method outperformed other resolution enhancement algorithms that depend on assumptions that can prove flawed.
“Our system learns various types of transformations that you cannot model because they are random in some sense, or very difficult to measure, enabling us to enhance microscopy images at a scale that is unprecedented,” Wang said.
Despite using an off-the-shelf computer — the equipment used in the study was similar to a standard gaming laptop — UCLA researchers were to produce super-resolution images in a fraction of a second. Rivenson said the system drastically simplifies super-resolution imaging and could readily be used by scientists without specialized expertise in imaging.
The study’s other authors are Yiyin Jin, Zhensong Wei, Ronald Gao and Harun Günaydin of UCLA; and Comert Kural of Ohio State University.
The research was supported by the National Science Foundation and the Howard Hughes Medical Institute. Imaging was performed at the Advanced Light Microscopy/Spectroscopy Laboratory at CNSI and at the Advanced Imaging Center at the HHMI Janelia Research Campus in Ashburn, Virginia.
http://newsroom.ucla.edu/releases/deep-learning-fluorescence-microscopy-super-resolution
